Quality trimming
In the first step, low-quality base calls are trimmed off from the 3' end of the reads before adapter removal. This efficiently removes poor quality portions of the reads.
The default Phred-score cutoff is 20 (-q 20). Trimming uses the BWA algorithm: the running sum of (Q - cutoff) is computed from the 3' end, and bases are removed up to the position with the maximum cumulative score.
Example: before and after
Section titled “Example: before and after”The plots below show a public dataset (DRR001650_1, Kobayashi et al., 2012) before and after trimming with -q 20.
| Before quality trimming | After quality trimming |
|---|---|
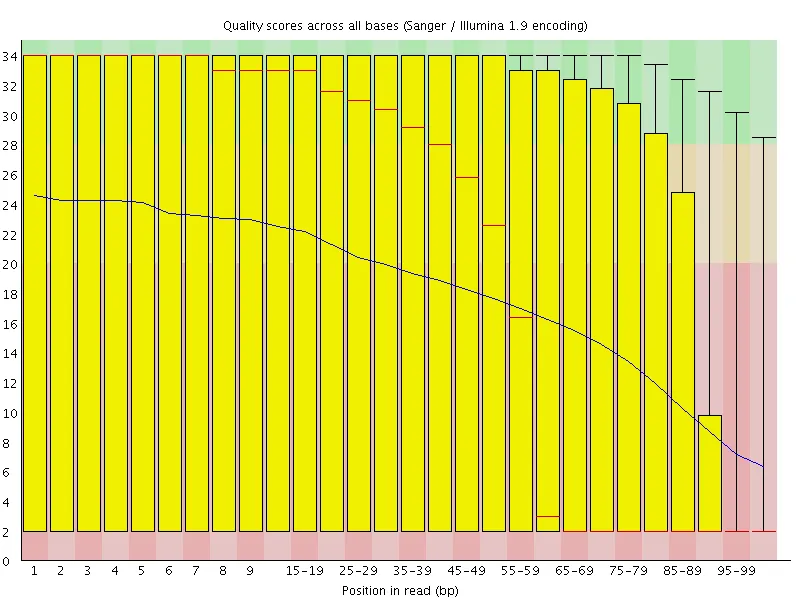 | 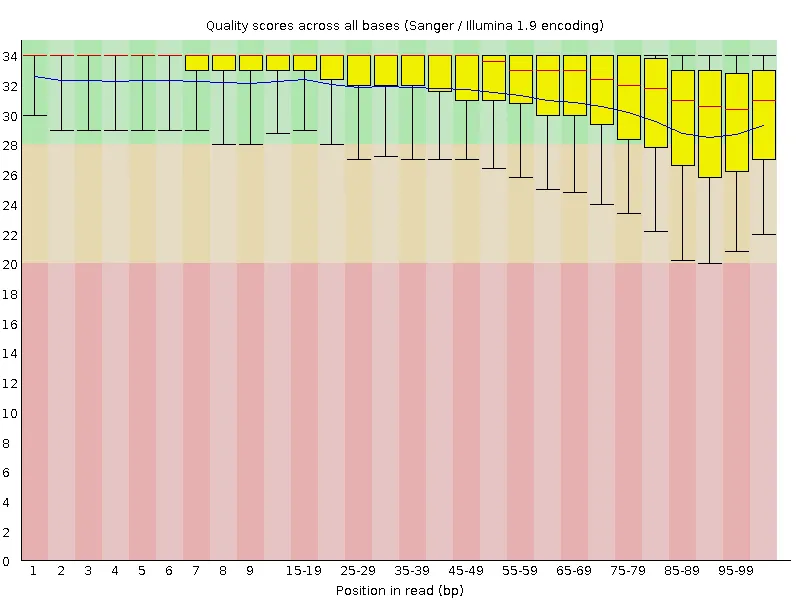 |
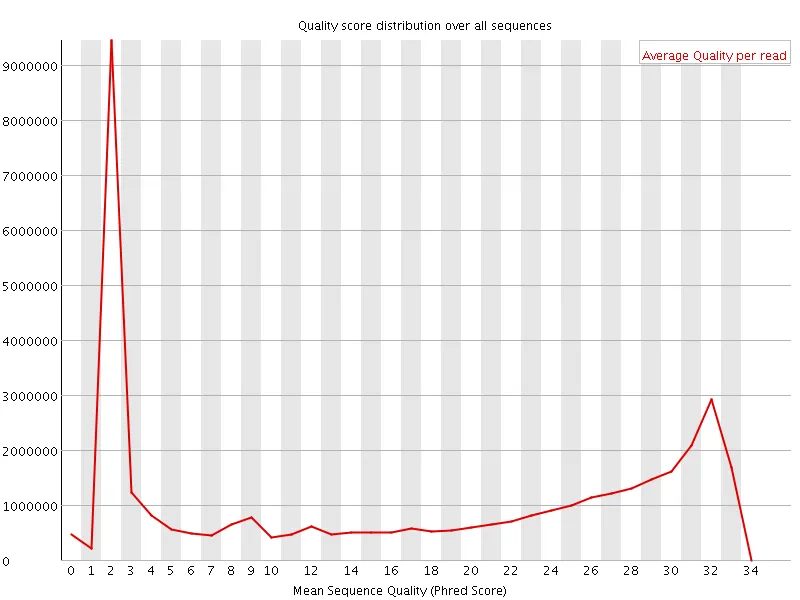 | 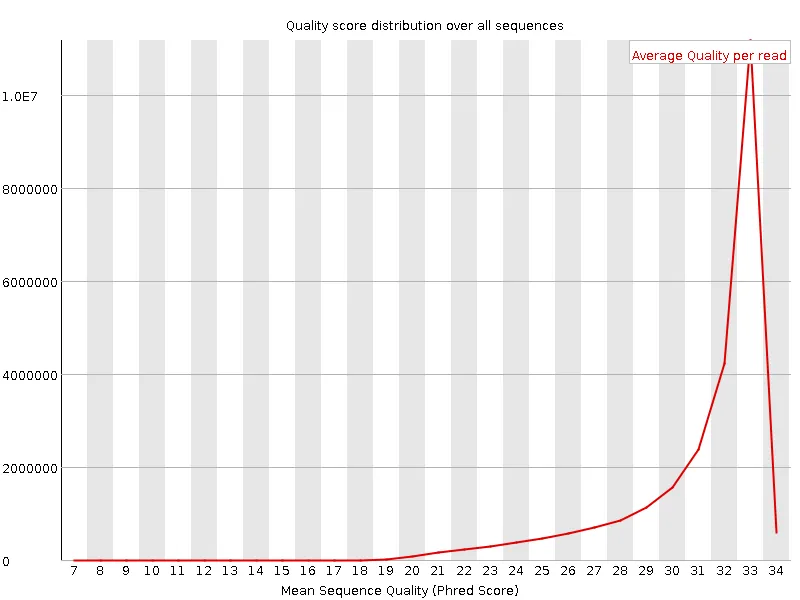 |
2-colour instrument quality trimming
Section titled “2-colour instrument quality trimming”Default -q quality trimming was designed for 4-colour Illumina chemistry (HiSeq, MiSeq). On 2-colour instruments (NextSeq, NovaSeq, NovaSeq X), no signal is encoded as G, which leads to spurious high-quality G-runs at the 3' end of reads with little signal. Pass --2colour N (or its alias --nextseq N) to use 2-colour-aware quality trimming, which treats trailing Gs as low-quality before applying the Phred cutoff.
trim_galore --2colour 20 input.fastq.gz--2colour and --nextseq are opt-in. They replace -q. They are independent of --poly_g, which is sequence-based and runs after quality trimming.
Phred encoding
Section titled “Phred encoding”Trim Galore defaults to Phred+33 encoding (Sanger / standard Illumina 1.8+). For very old data in Phred+64, pass --phred64.
What's logged
Section titled “What's logged”The chosen cutoff and encoding are recorded in the trimming report:
Quality Phred score cutoff: 20Quality encoding type selected: ASCII+33The Cutadapt-compatible block in the same report shows the total bases trimmed by quality.
Related flags
Section titled “Related flags”-q INT. Phred quality cutoff (default 20).--phred33/--phred64. Phred encoding (default Phred+33).--2colour N/--nextseq N. Opt-in 2-colour-aware trimming, replaces-q.